The potential for Oxford Nanopore’s MinION to revolutionise genomics has been well documented, and for good reason. As we and others have mentioned before, they have drastically reduced the size of sequencers so that their device will fit in a pocket. Perhaps as importantly, the protocol to prepare samples for sequencing is significantly simpler than traditional sequencing, raising the possibility of this technology being used by more and more people interested in genetics - including us!
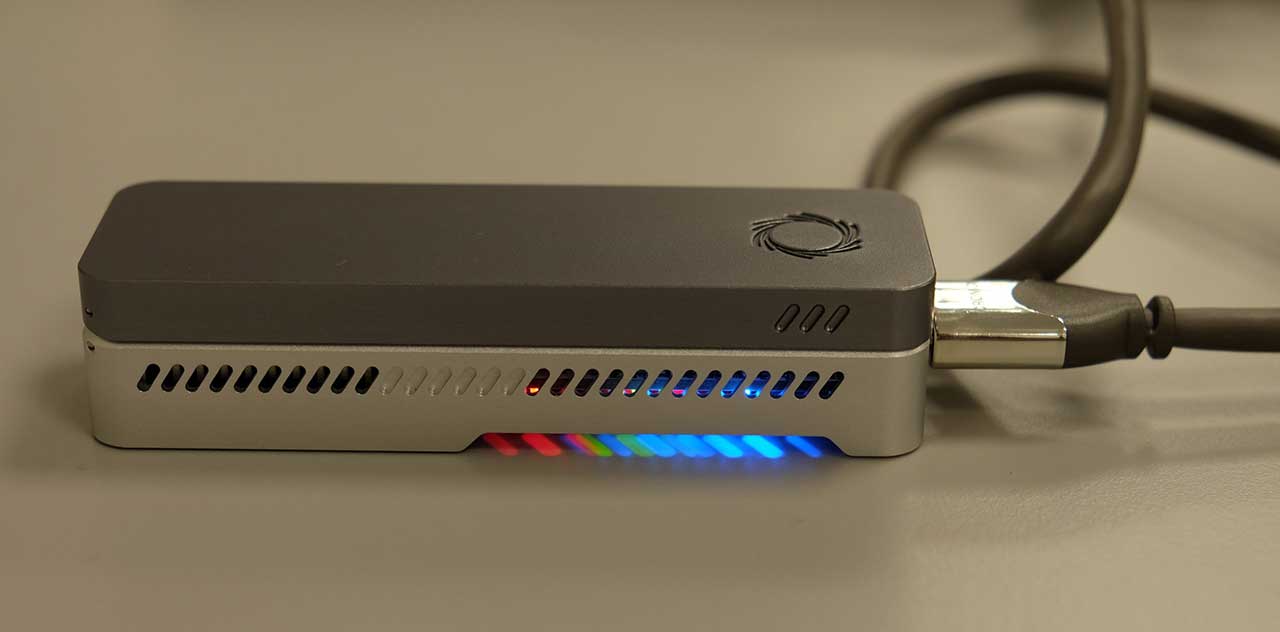

Our MinION running
Having recently conducted a burn-in experiment to check that our MinION works and to familiarise ourselves with the protocol, we had been racking our brains for a good experiment to use our remaining 2 flow cells for.
Sequencing for soil health
As one of our current focuses is soil health we thought it would be interesting to investigate whether we could extract DNA from the organisms found in two different soil samples, sequence them using MinION, and use the species identified to infer information about the soil properties. After some further digging on Google we were unable to find evidence of anyone else who had tried this, giving us even more reason to give it a go!
The analysis we were intending to do was made possible thanks to an app for Metrichor (Oxford Nanopore’s analysis software) called What’s In My Pot (WIMP). WIMP connects to the NCBI database of bacterial, fungal and viral genomes and matches reads sequenced by MinION to their respective species.
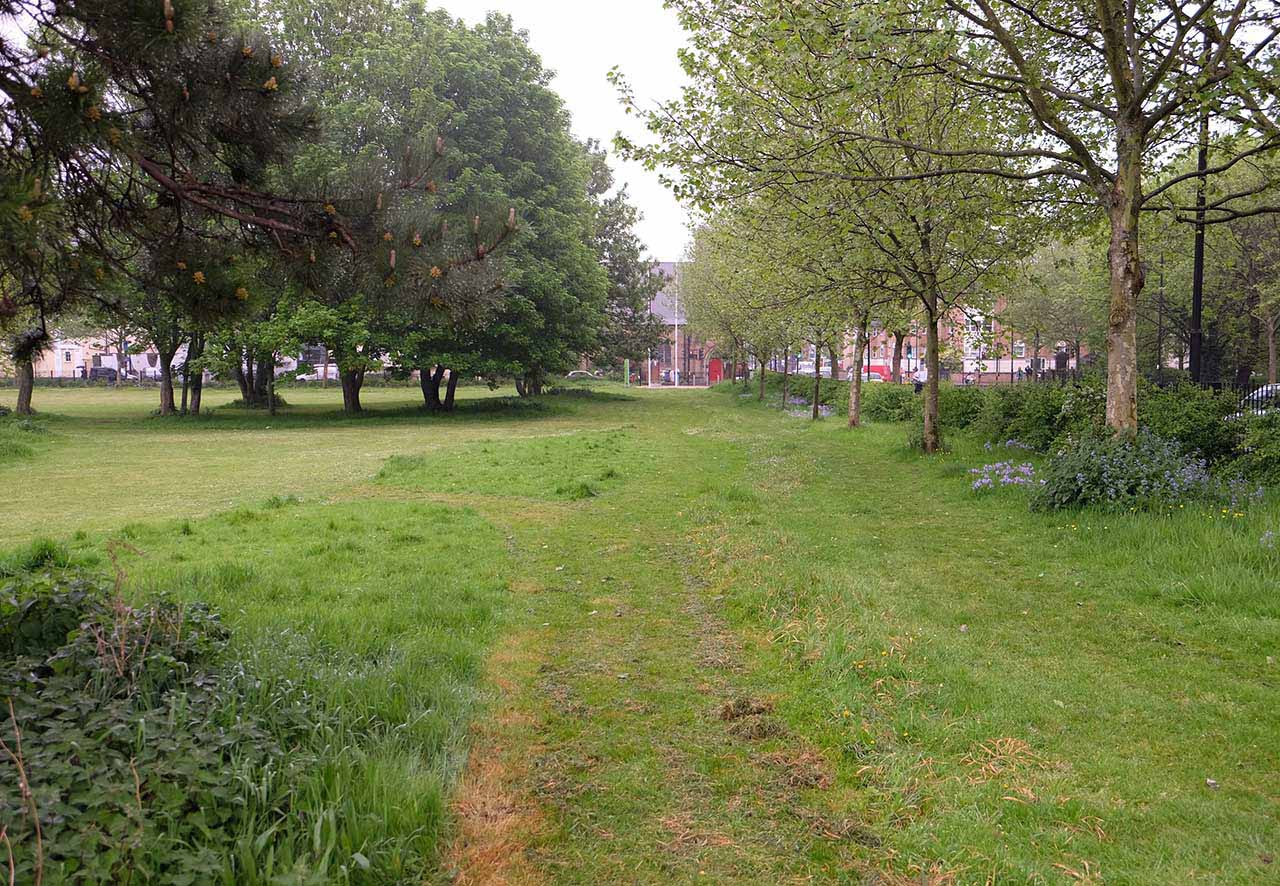
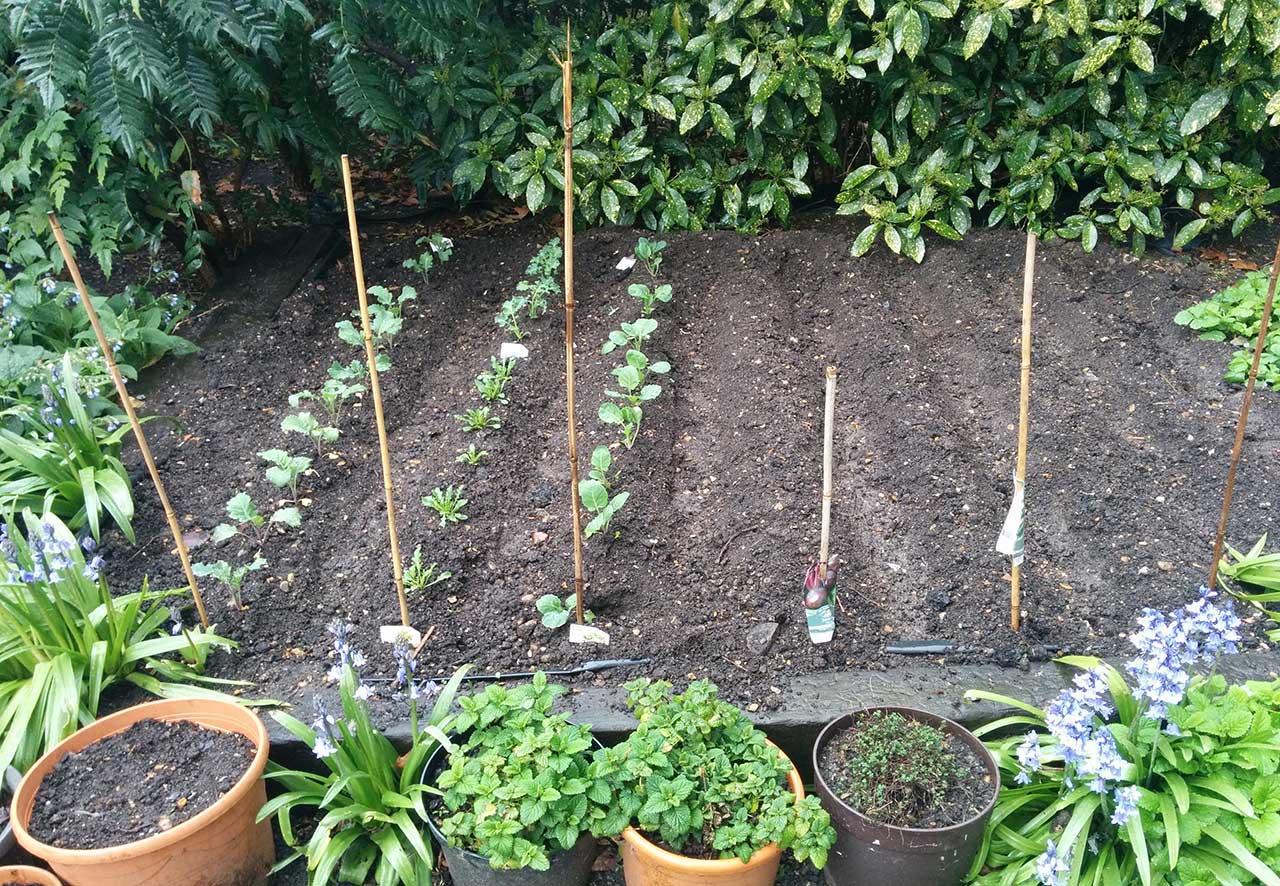


Mile End Park and a Bloomsbury vegetable patch were our chosen places to take soil samples
Our first steps were to identify two places to collect soil samples from, and to find a way to extract DNA from the bacteria and fungi found within them (known as the metagenome). For the soil we settled on two samples from London: one from a worn patch of ground in Mile End Park, and one from a shared vegetable patch. Our hypothesis was that there may be detectable differences between the cultivated and non-cultivated soils.
Extracting DNA from soil
DNA extraction was easier than you may expect. We were able to order a DNA extraction kit from MO BIO called Powersoil which was simple to use and took about two hours to go from sample to DNA in a lab at Imperial College London. Ordinarily a biomedical lab, we raised a few eyebrows with our soil experiments!
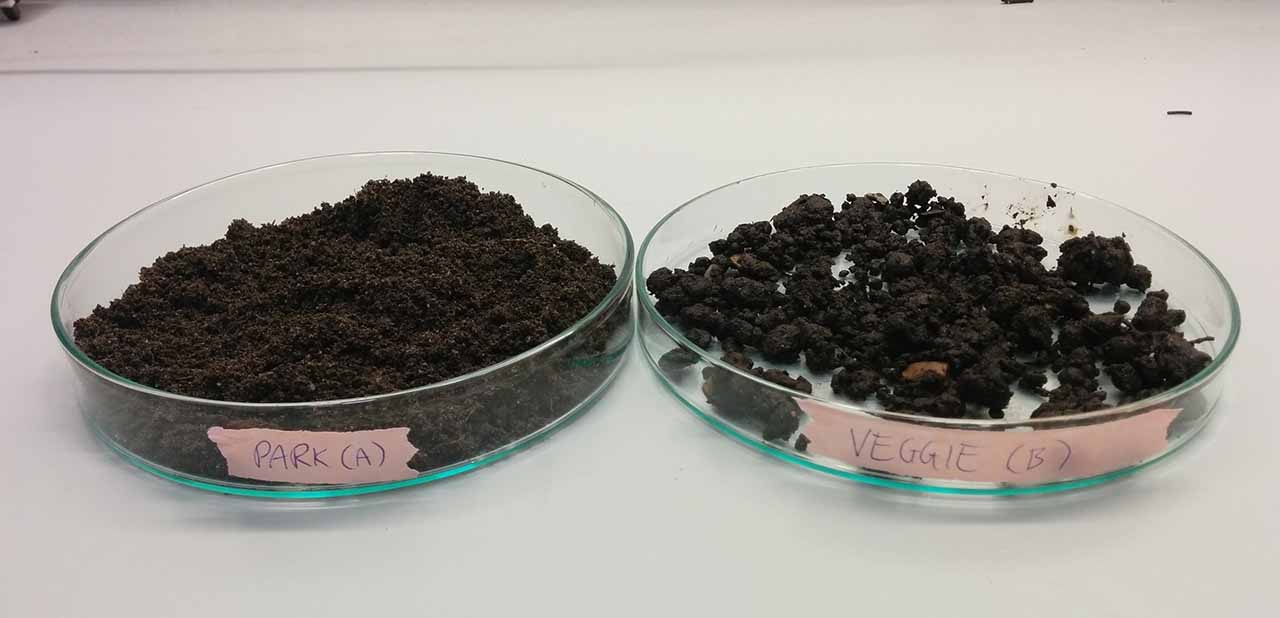

Our soil samples ready for DNA extraction
The next step was to find somewhere to prepare the DNA and to run the MinION sequencing. As we were well aware from our burn-in experiment, there are a number of pieces of lab equipment that are necessary that we do not have access to, and so we needed to find someone with lab space who would be willing to work with us on this mini-project.
Visiting York
Through twitter we managed to find James Chong at The University of York, who had recently given a talk on using MinION to sequence bacterial 16s ribosomal RNA, and asked if he might be interested. Thankfully for us, James was incredibly generous with his time and knowledge, introduced us to his equally helpful post-doc Anna Alessi who had been using MinION, and offered to have us visit his lab and run the experiment.
Excited to be in @insanity_one lab today @yorkuniversity! sequencing soil creatures with the #minion @nanopore pic.twitter.com/3KbyjYRoVR
— Science Practice (@sciencepractice) 5 May 2016
When we arrived in York we were warmly welcomed by James, Anna and Farah Shahi who is a medical doctor at the start of a research project using MinION and is learning lab techniques. After picking James’s brain for advice for our soil health projects we quickly got going on the MinION protocol, starting by checking whether we had enough DNA using Qubit, a fluorometer.
We were relieved to find that we had more than enough DNA, and so proceeded with the same protocol as we did for the burn-in, only this time we prepared two samples in parallel: the sample from the park and the vegetable patch. By the end of the protocol we were pleased to have two good DNA libraries that met Oxford Nanopore’s suggested quantity requirements.
Relieved to find that we extracted enough #DNA from soil to sequence on the @nanopore #minion today #dnaparty pic.twitter.com/ZnMWaqXI0G
— Science Practice (@sciencepractice) 5 May 2016
While the wet-lab component of the day went without problem, we did have some issues with running MinION itself. Firstly we were unable to get our laptop to recognise the MinION. We were aware that this could be temperamental from our experiences in the burn-in, however for this experiment it simply did not manage to acknowledge the device. Fortunately for us, James has five MinIONs permanently connected to five laptops and so we were able to use his.
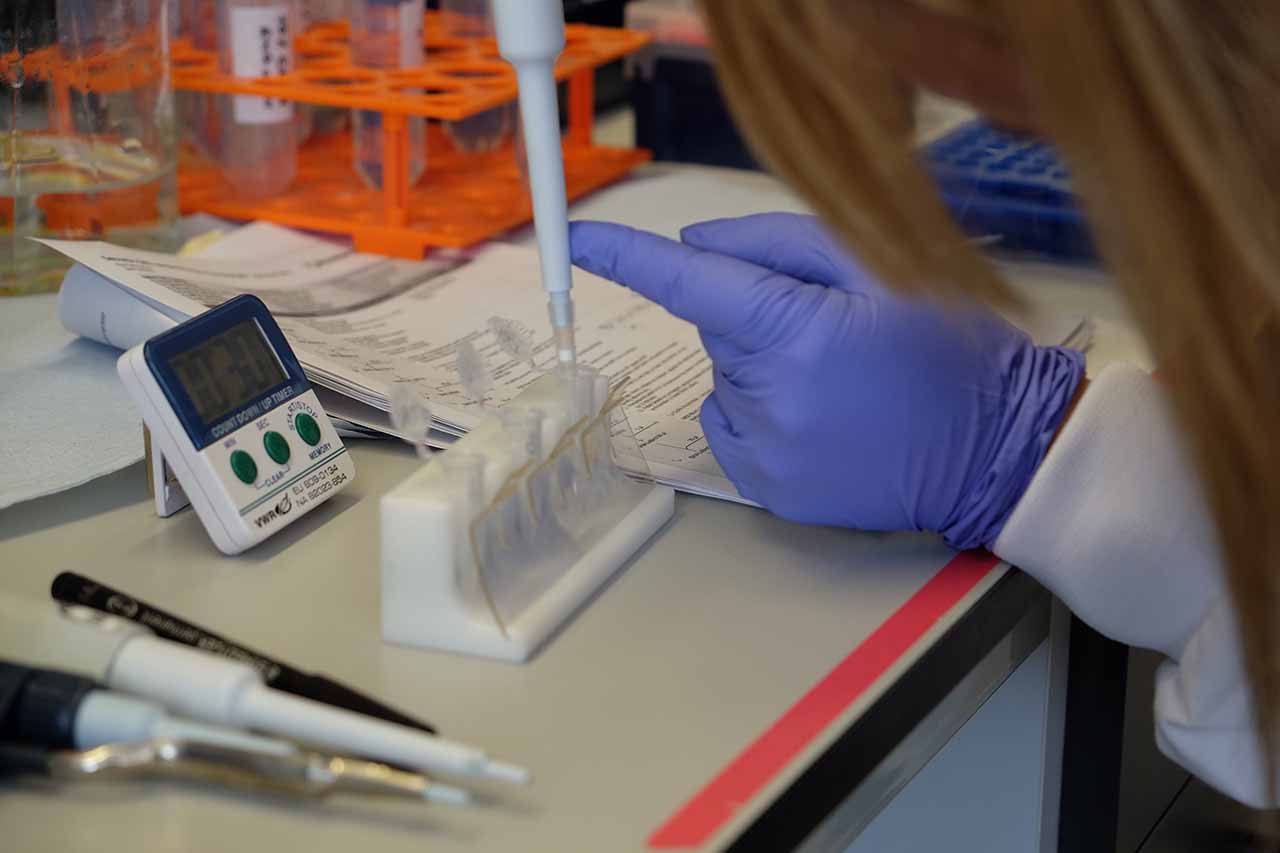

Using DNA binding magnetic beads and a magnetic tube rack to isolate our DNA samples
Silent flow cells
Having decided to use James’s MinION we then tested out our two flow cells. Although we had stored them how MinION recommended, and both were visually in good condition, the test run indicated that we had 0 working pores on either flow cell. Despite consulting Anna, a bioinformatician and even the Head of Genomics, we were not able to get to the bottom of this. We suspect there was a serious underlying problem with these flow cells, rather than a degradation of pores.
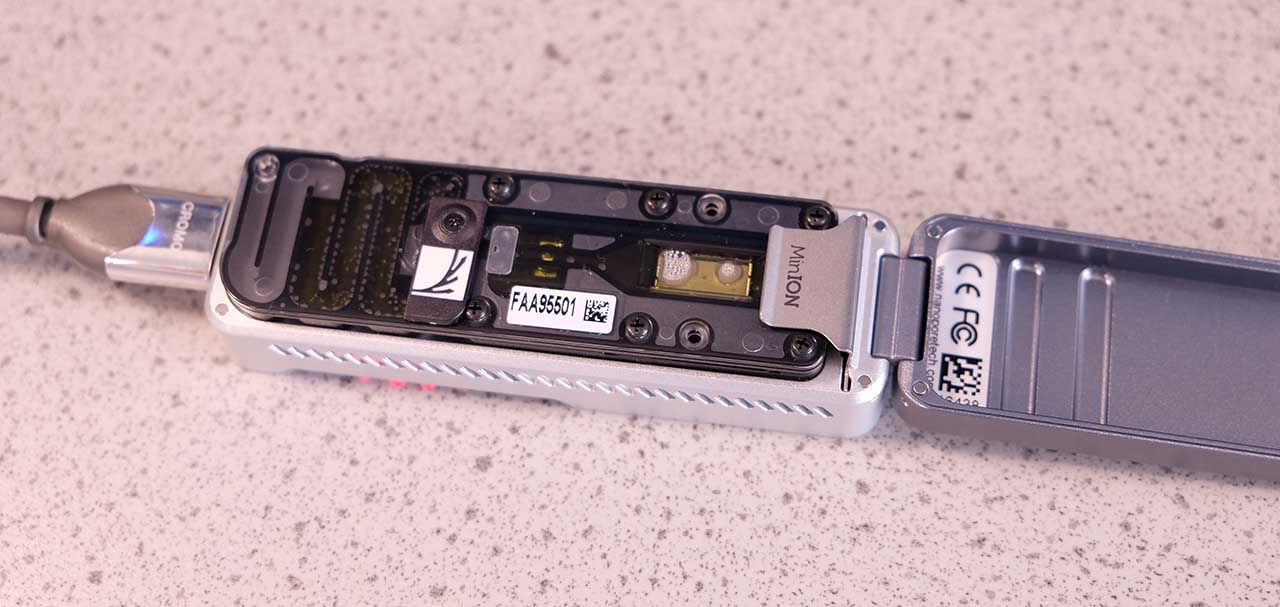

A MinION flowcell, the bubbles that are visible can hinder the experiment
Fortunately Anna and James came to the rescue again, and allowed us to use two of their spare flow cells. Although they were not in excellent condition (there were visible bubbles covering the pores), and only had around 150-400 active pores, after getting this far with our experiment it would have been a shame not to finish.
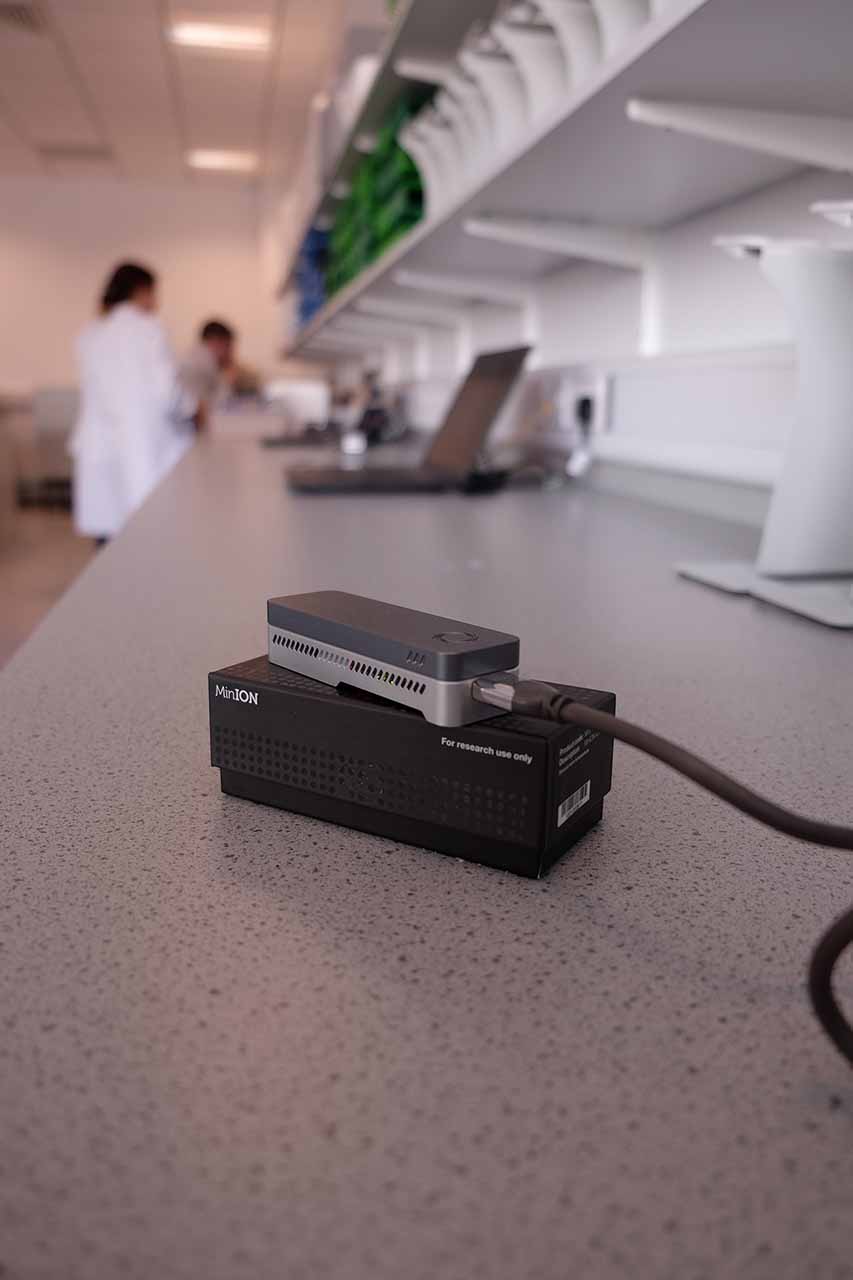


At work in the lab setting up our MinION runs
We ran the MinION overnight, and in the morning, reloaded with more of the libraries we had prepared and ran again (this is a trick that James and Anna have found improves the number of usable reads). As expected the data we got was relatively poor quality, with the majority of reads not meeting Metrichor’s quality control.
Genetic sequencer may be pocket-sized but we have the entire teaching lab to ourselves to run it @nanopore #minion pic.twitter.com/q0oQEAlb4u
— Science Practice (@sciencepractice) 6 May 2016
What’s in my pot?
Despite having few good quality reads we decided to run WIMP for both samples, to see what it was able to identify in each sample, and to get a better idea of how the app works. Although we can’t draw any confident conclusions from what we found, there were some small hints that it would be possible to use MinION combined with WIMP to gather useful information about soil.
For example several bacteria capable of metabolising common environmental pollutants were identified in the sample of soil from the park (Nocardioides, V. paradoxus), while the soil from the vegetable patch, which has had manure and compost added to it, had bacteria often found in sewer sludge and wet soil (Thauera). Both samples had many different bacteria involved in the nitrogen cycle.
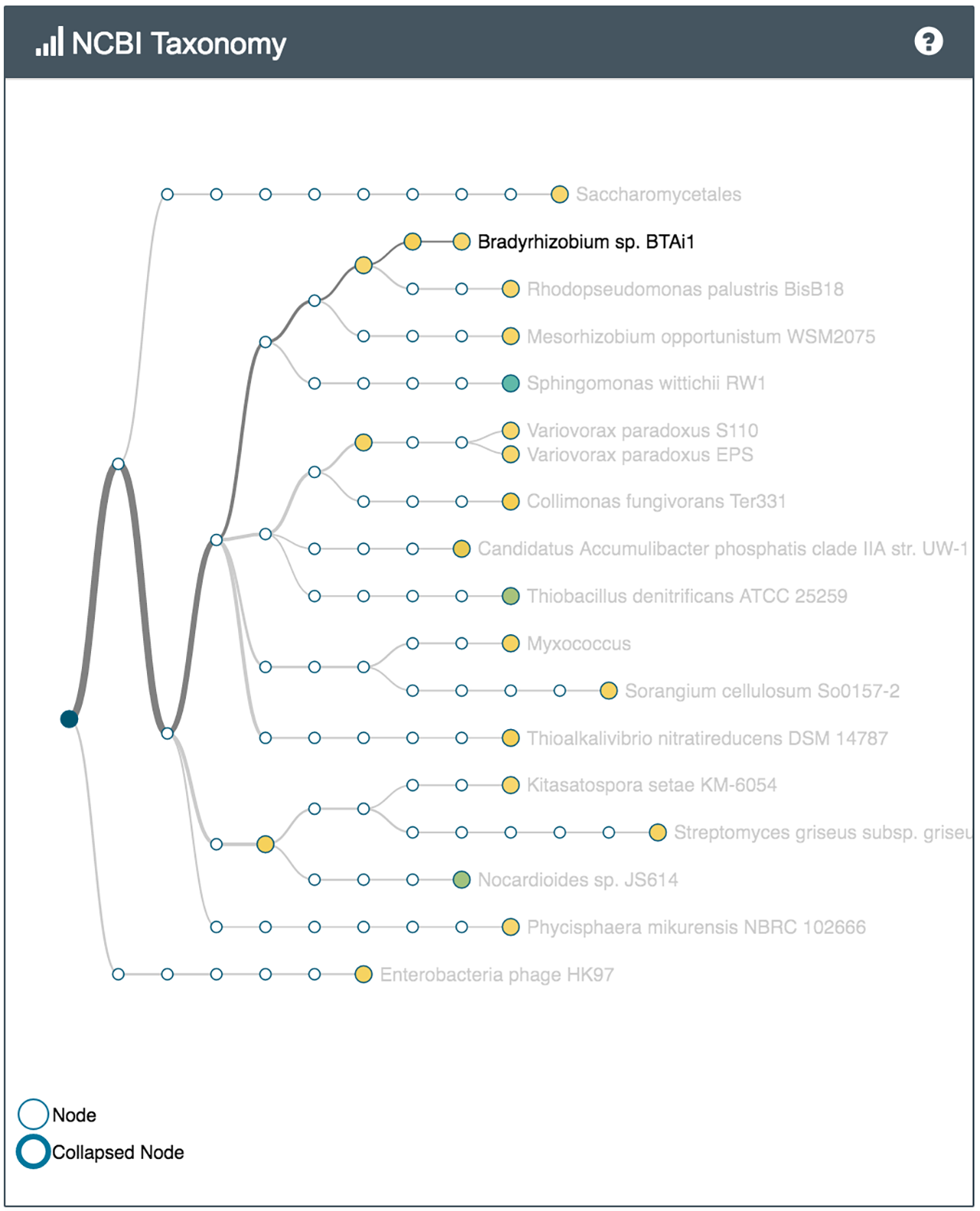
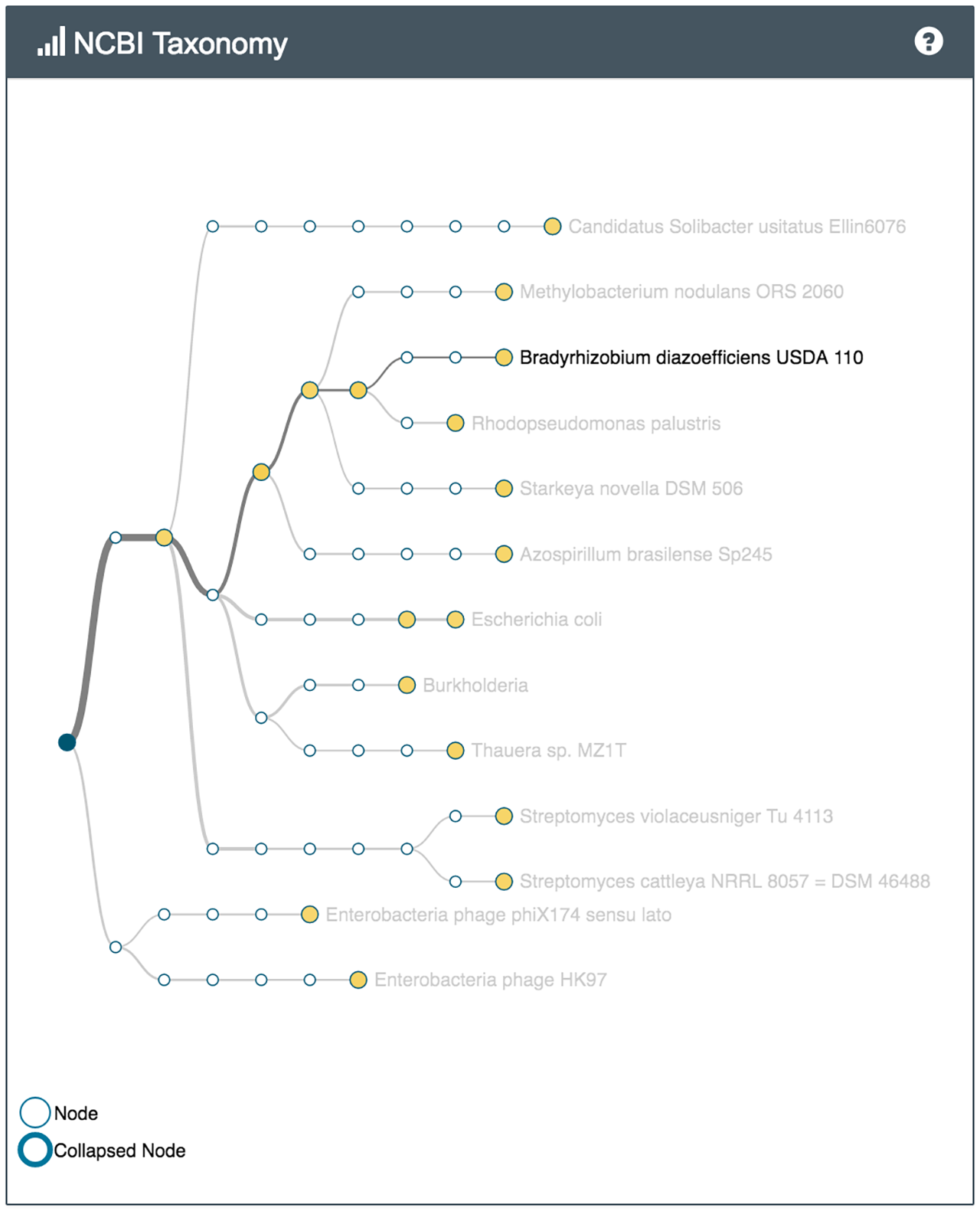
What's in my pot showing some of the species identified in the park sample (left) and the vegetable patch sample (right)
Cautious optimism
It is important not to over-analyse what we saw - the low read count meant that anything we didn’t see is as likely to have not been sequenced as absent in the sample, and anything we did see could easily be misinterpreted by the software as the wrong species. Also these were the results of just two samples. The entire experiment would need to be replicated in order to begin to make any kind of solid conclusions about inferring soil properties from their metagenomes using this method.
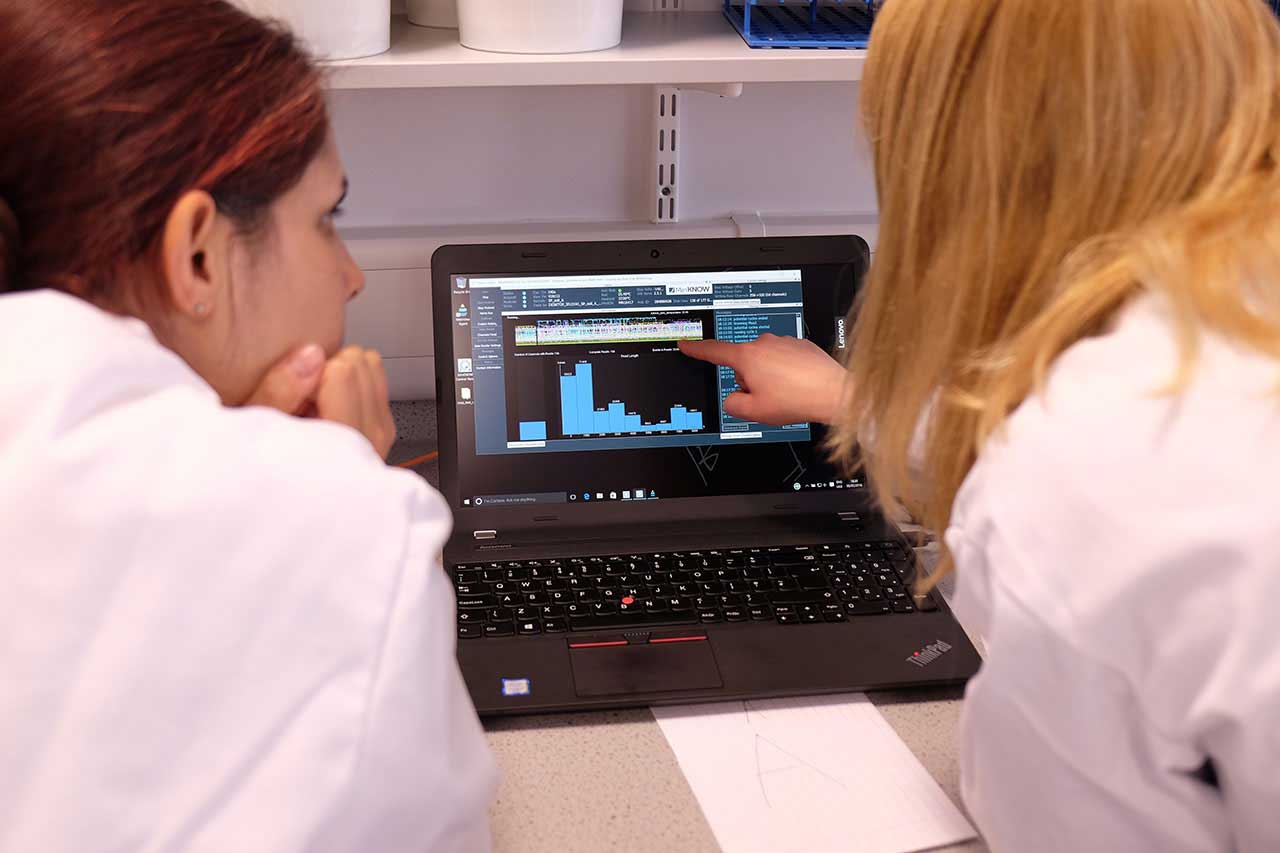

Watching our data come in on the MinION desktop software MinKNOW
Nevertheless the experiment can give broad encouragement that it should be possible to gain a rich understanding of a soil metagenome, and from this information infer properties of soil health and environment using MinION and WIMP. A distinct advantage of using MinION is that, with good quality data, you can make estimates about the proportions of different species in a sample, as this is related to the number of reads for each species.
Although there are still many technical challenges with using MinION, with lab space, some patience and guidance from someone with experience, it is a technology with remarkable potential. While it would take a lot of effort and significantly better microbial knowledge than we currently possess, the possibility of using nanopore sequencing to gain a better understanding of soil health is genuinely exciting.
If you are interested in discussing this project, our results or having a look at the data we collected, please get in touch!

