We worked with a genomics startup to enable the users of their variant identification platform to understand and build trust in the prioritisation of disease causing variants.
We designed a bespoke data visualisation that provided a peek inside the ‘black box’ of a bioinformatics algorithm.
We worked with Repositive on their rapid journey to build a new platform to help cancer researchers find experimental models.
We worked from concept to alpha release, providing user interface design and user research support.
We designed a novel way to quickly understand information about Single Nucleotide Polymorphisms (SNPs).
SNPshot draws on principles of information hierarchy to help distill and display key characteristics of SNPs as part of decision-making processes in clinical genetic.
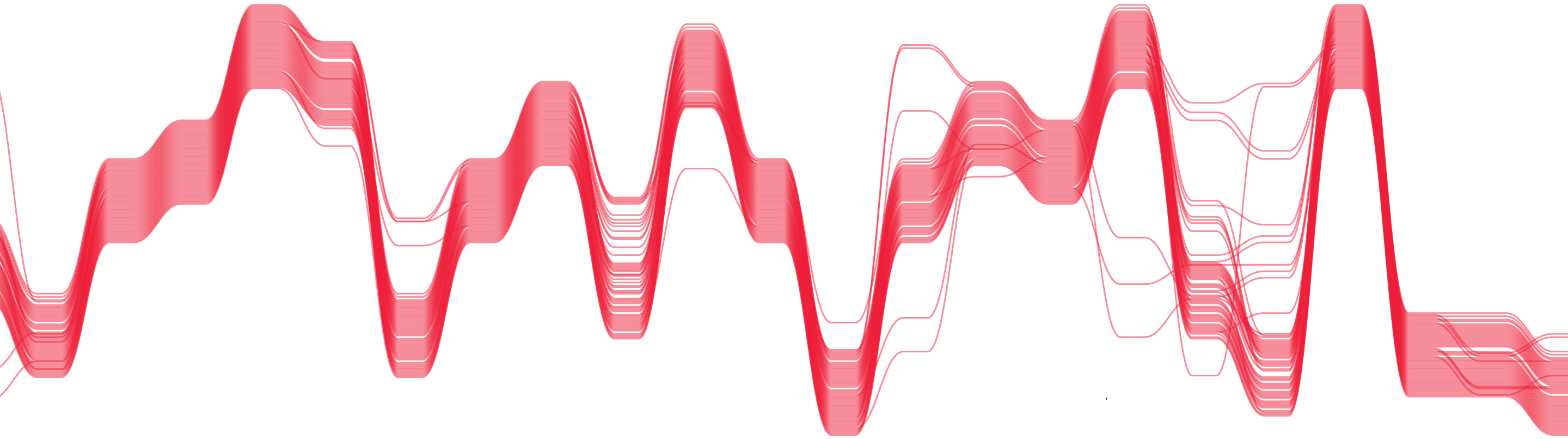
We designed and developed a new way of visualising multiple sequence alignment data - Sequence Bundles.
Sequence bundles encourages exploration of multiple sequence alignment data, and preserves continuity of information, allowing the discovery of patterns in data that could otherwise remain hidden.
Exploring the possibility of using MinION genetic sequencer to measure the microbiological biodiversity of soil - without our own lab.
We tested the usability of hardware and software for sequencing, and found insights into how agriculture could use new technology to monitor soil health.